HSA: integrating multi-track Hi-C data for genome-scale

HSA: integrating multi-track Hi-C data for genome-scale

Hi-C analysis: from data generation to integration

HiC-bench workflow. Raw reads (input fastq files) are aligned and then

ParticleChromo3D: a Particle Swarm Optimization algorithm for chromosome 3D structure prediction from Hi-C data, BioData Mining
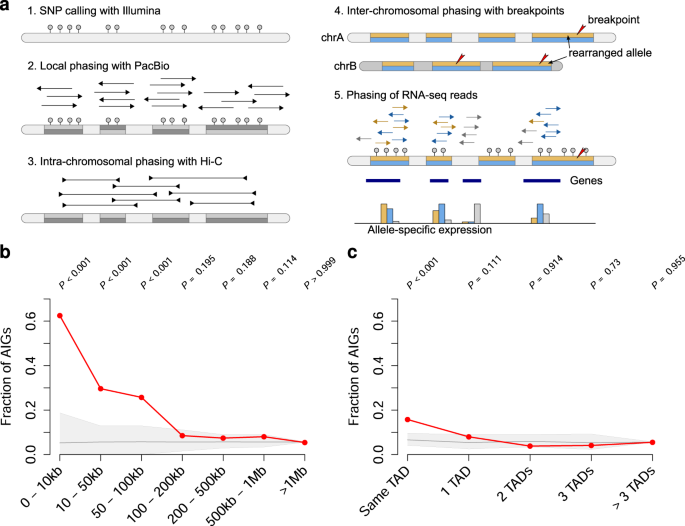
Integration of Hi-C with short and long-read genome sequencing reveals the structure of germline rearranged genomes
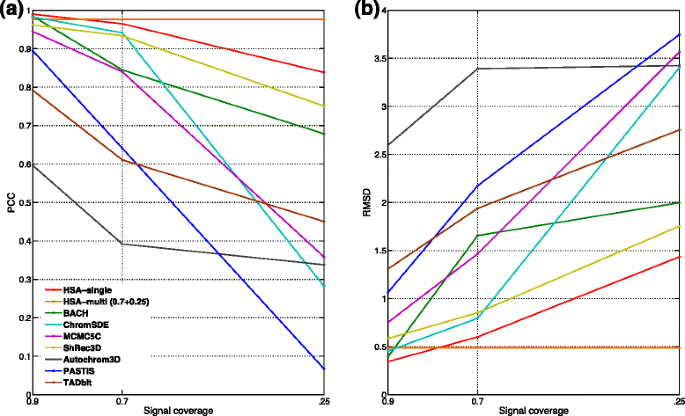
HSA: integrating multi-track Hi-C data for genome-scale reconstruction of 3D chromatin structure, Genome Biology

HSA: integrating multi-track Hi-C data for genome-scale reconstruction of 3D chromatin structure, Genome Biology

HSA: integrating multi-track Hi-C data for genome-scale reconstruction of 3D chromatin structure
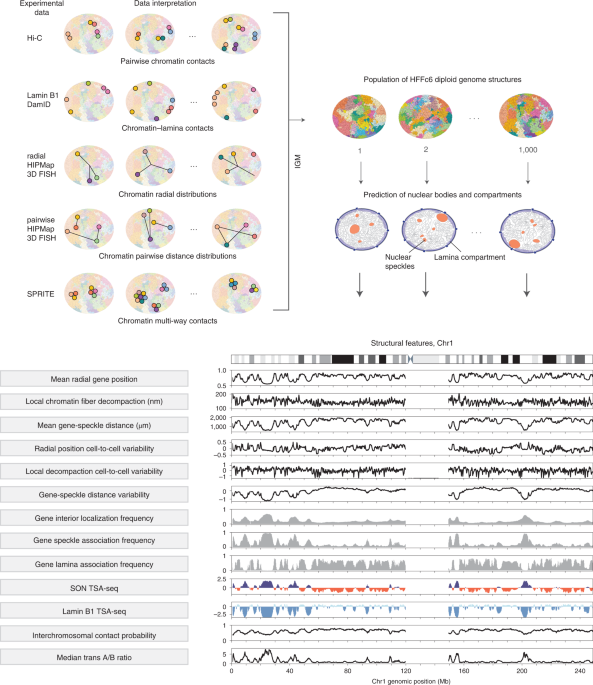
Integrative genome modeling platform reveals essentiality of rare contact events in 3D genome organizations

Sci-Hi-C: a single-cell Hi-C method for mapping 3D genome organization in large number of single cells

VEHiCLE: a Variationally Encoded Hi-C Loss Enhancement algorithm for improving and generating Hi-C data

Personal Omics Profiling Reveals Dynamic Molecular and Medical Phenotypes: Cell
Integrating Hi-C links with assembly graphs for chromosome-scale assembly

An Overview of Genome Organization and How We Got There: from FISH to Hi-C


