Invitrogen™ ERCC ExFold RNA Spike-In Mixes

Invitrogen™ ERCC ExFold RNA Spike-In Mixes
ERCC ExFold RNA Spike-In Mixes provide a set of external RNA controls that enable performance assessment of a variety of technology platforms used for

LaboShop, Products
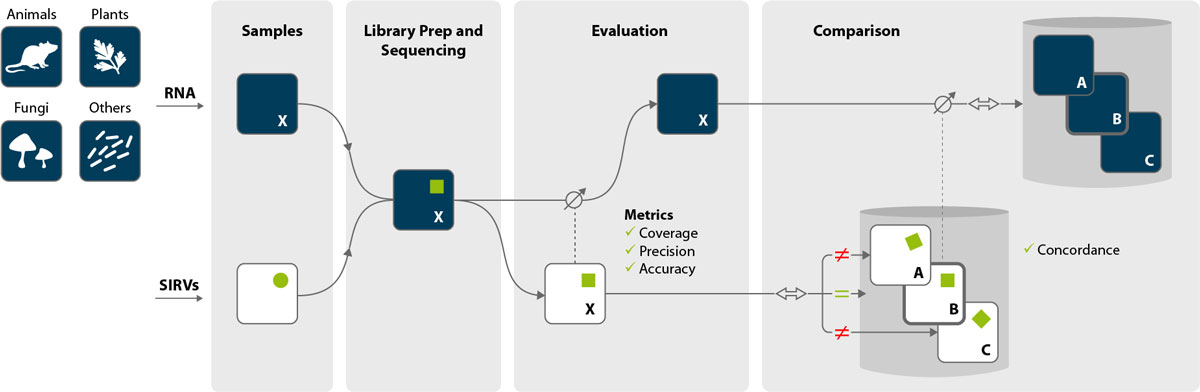
SIRVs (Spike-in RNA Variant Control Mixes)

LaboShop, Products

High-throughput full-length single-cell RNA-seq automation

Spaceflight influences gene expression, photoreceptor integrity, and oxidative stress-related damage in the murine retina
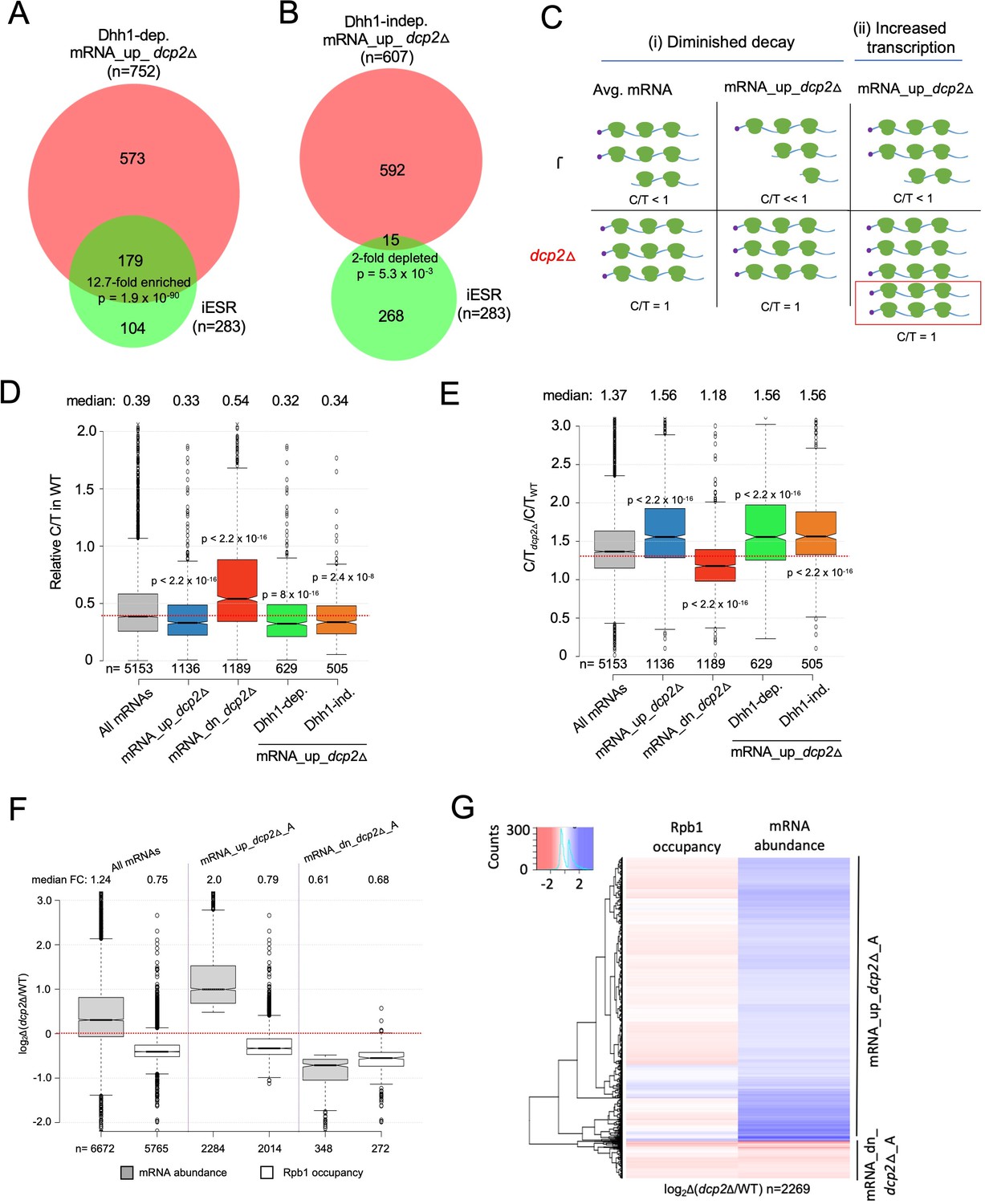
Decapping factor Dcp2 controls mRNA abundance and translation to adjust metabolism and filamentation to nutrient availability
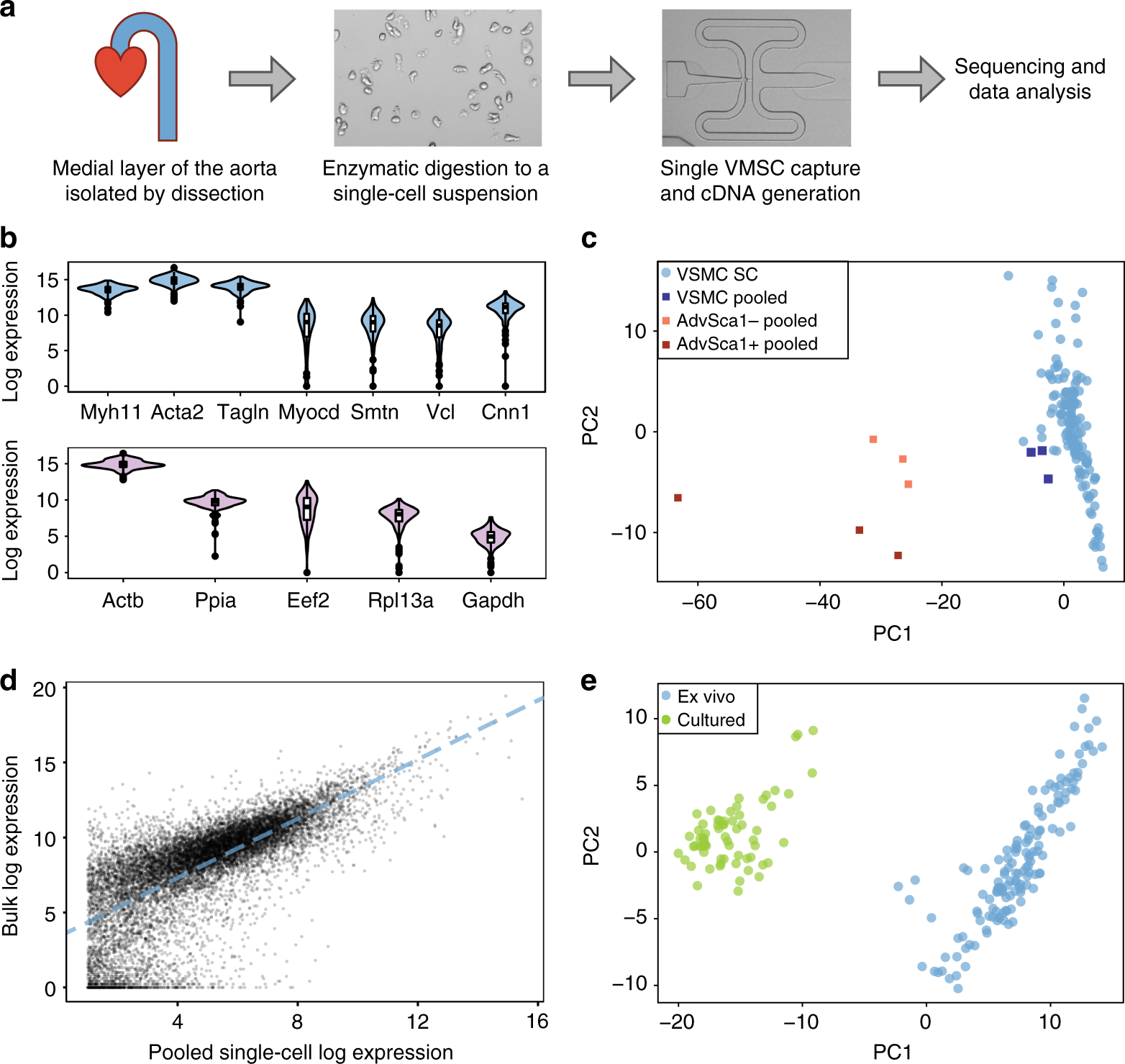
Disease-relevant transcriptional signatures identified in individual smooth muscle cells from healthy mouse vessels

Single-cell RNA sequencing reveals homogeneous transcriptome patterns and low variance in a suspension CHO-K1 and an adherent HEK293FT cell line in culture conditions - ScienceDirect
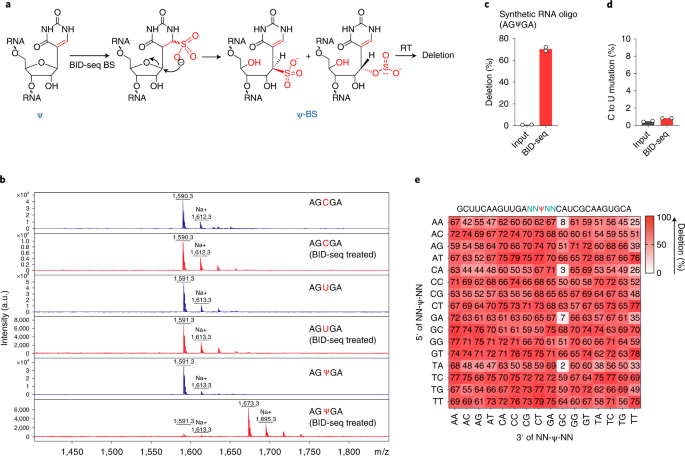
Quantitative sequencing using BID-seq uncovers abundant pseudouridines in mammalian mRNA at base resolution

The Role of Spike-In Standards in the Normalization of RNA-seq

Evaluation of the External RNA Controls Consortium (ERCC) reference material using a modified Latin square design, BMC Biotechnology

SARS-CoV-2 Nucleocapsid/Spike Protein (RBD) Recombinant Protein (RP-87706)

Single-cell RNA sequencing reveals homogeneous transcriptome patterns and low variance in a suspension CHO-K1 and an adherent HEK293FT cell line in culture conditions - ScienceDirect

The Role of Spike-In Standards in the Normalization of RNA-seq

Revisiting Global Gene Expression Analysis: Cell


